-
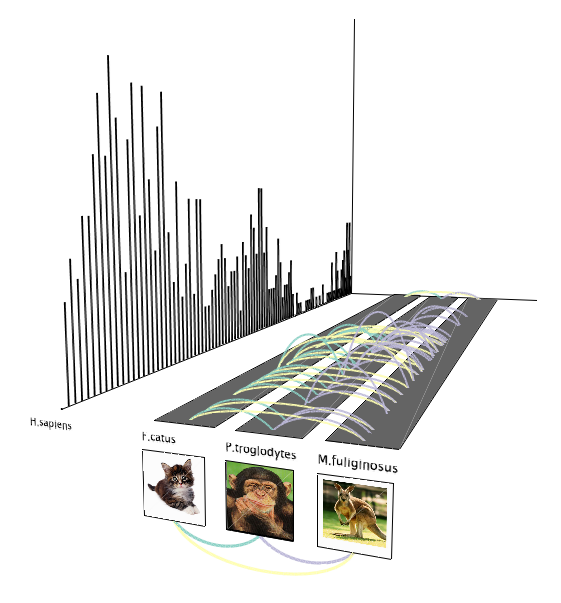 Conserved transcription events
Conserved transcription eventsPeaks of transcription throughout the cell cycle are visualized for human (bar chart) and connections between 3 other species are shown if they share conserved genes that have the transcription peak at the corresponding time point.
-
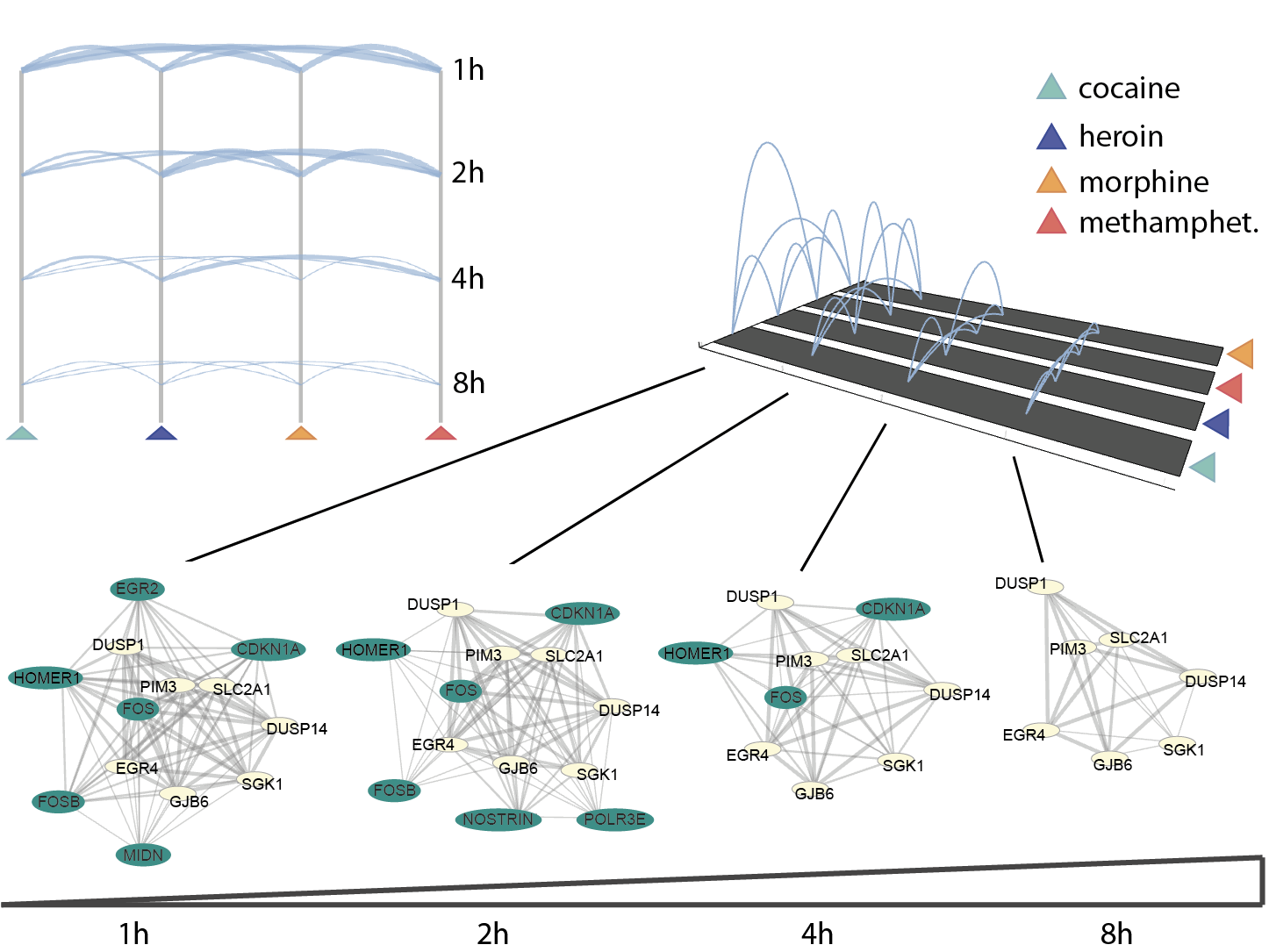 Similar mechanisms of drug action
Similar mechanisms of drug actionThe number of commonly highly expressed genes upon induction of different drugs is displayed through drug connections at every time point. Changes in the network of genes sharing drug phenotypes are also shown through time.
-
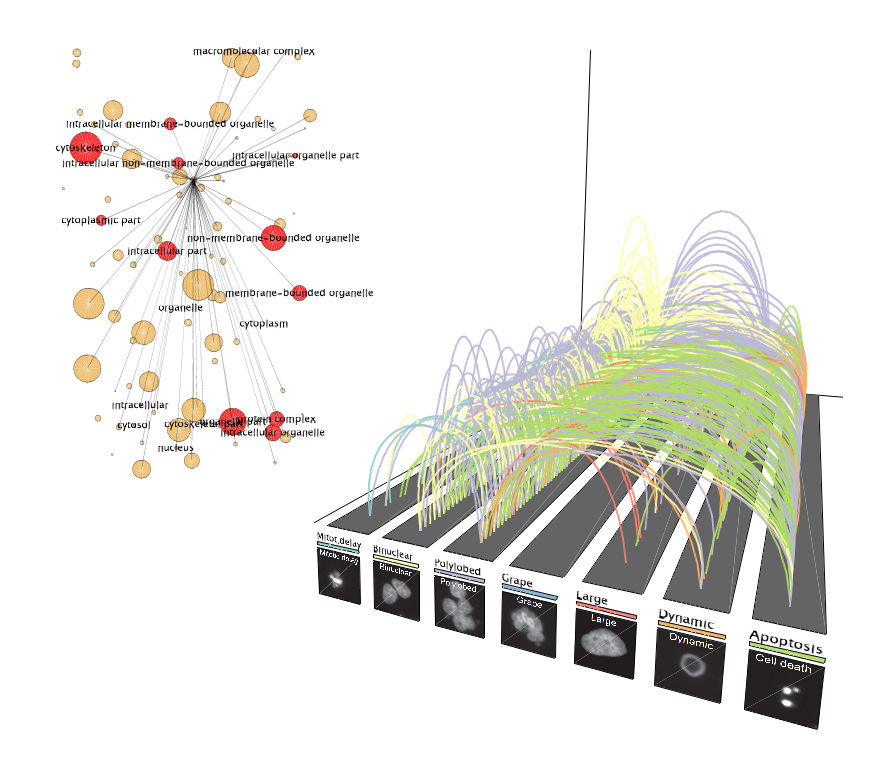 Phenotypic transitions in the cell cycle
Phenotypic transitions in the cell cyclePhenotypic transitions of cell populations upon knockdown of cell cycle genes are visualized using arcs to connect phenotypes. The GO network dynamically highlights functions of the genes responsible for the transitions at one time point.
-
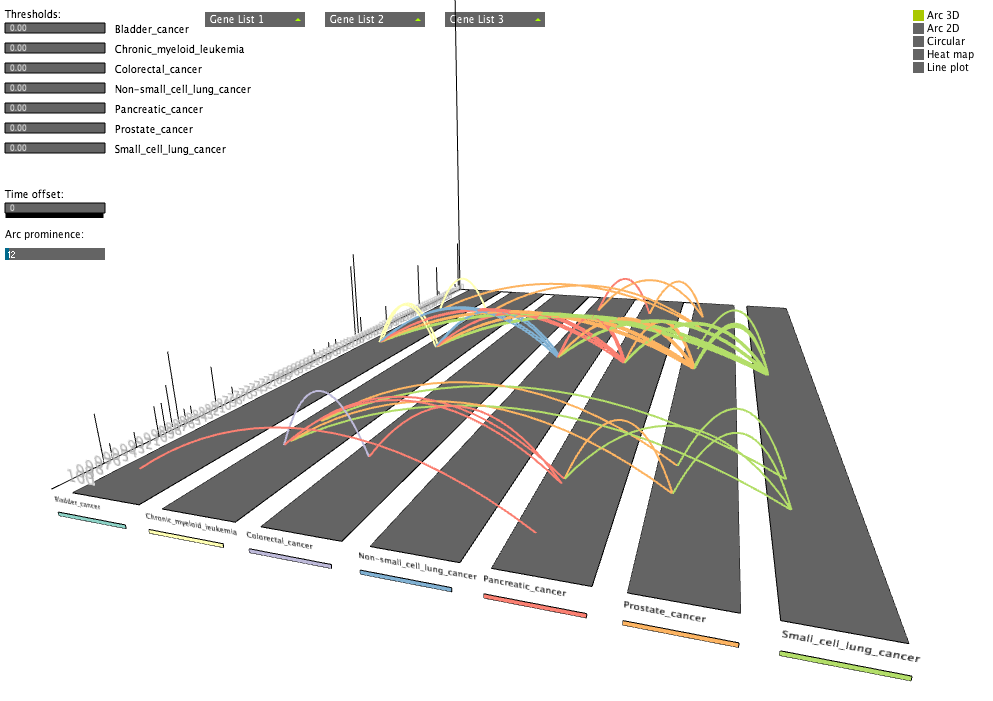 Transcription events linking diseases
Transcription events linking diseasesVisualizing peaks of transcription throughout the cell cycle and the corresponding cancer pathways that are pairwise enriched for the corresponding genes.
-
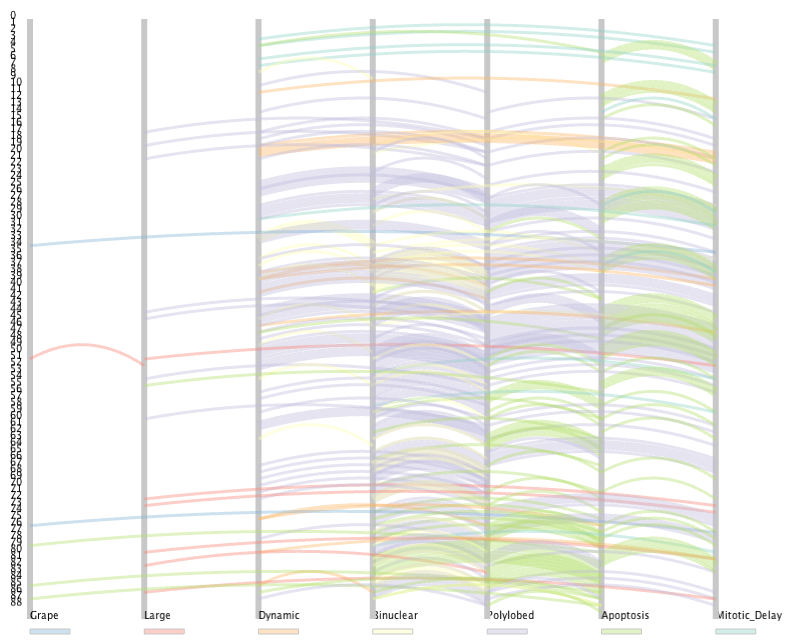 Phenotypic transitions in 2D
Phenotypic transitions in 2DPhenotypic transitions of cells upon single perturbation events are visualized in 2D arc mode. Transparency of arcs can be modified for optimal display.
-
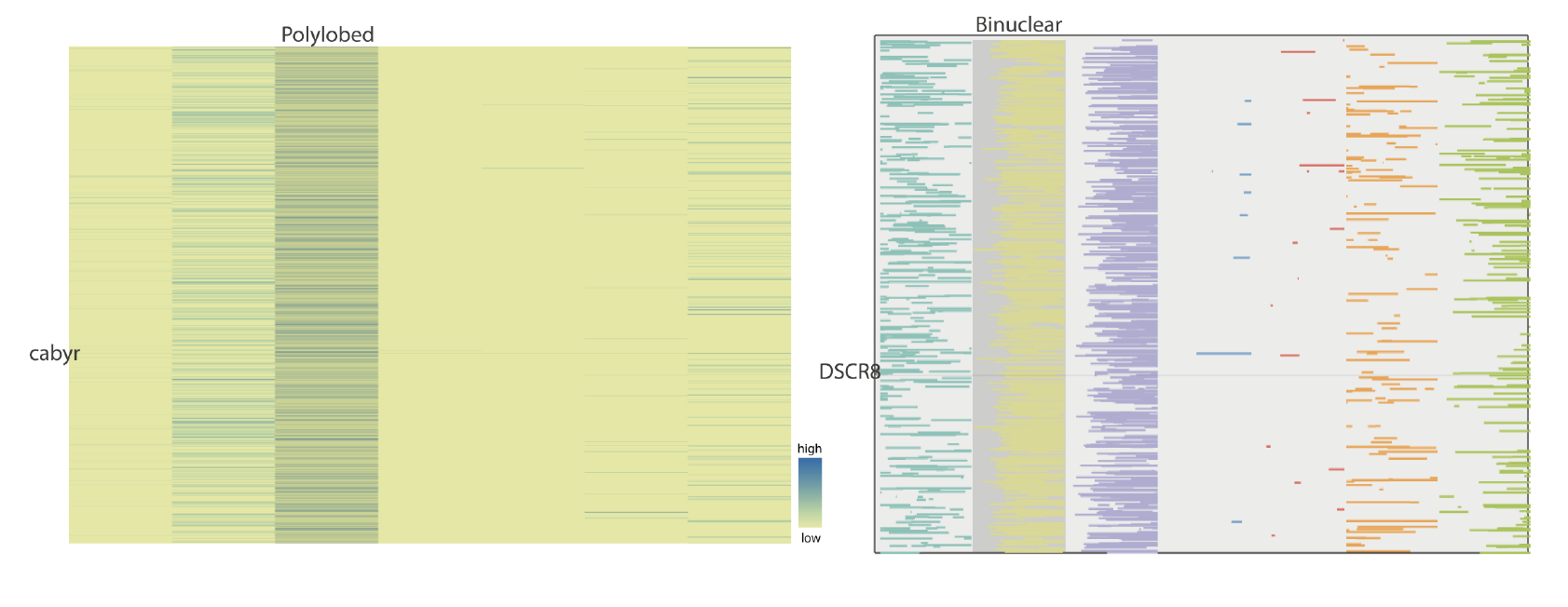 Heat maps and line plots
Heat maps and line plotsHeat maps and line plots of gene-associated values for each phenotype are interactive, highlighting gene and phenotype name upon hovering over with the mouse.
-
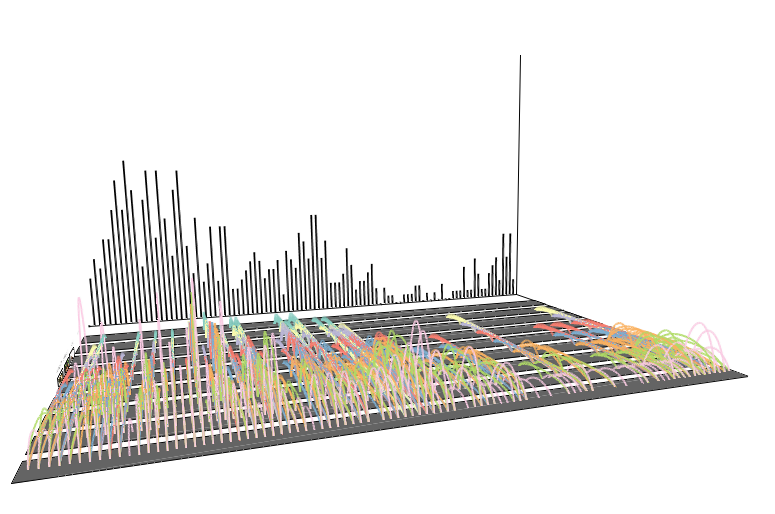 Perspectives: arc mode
Perspectives: arc modeArc mode displays connections between species that share conserved transcription events throughout the cell cycle.
-
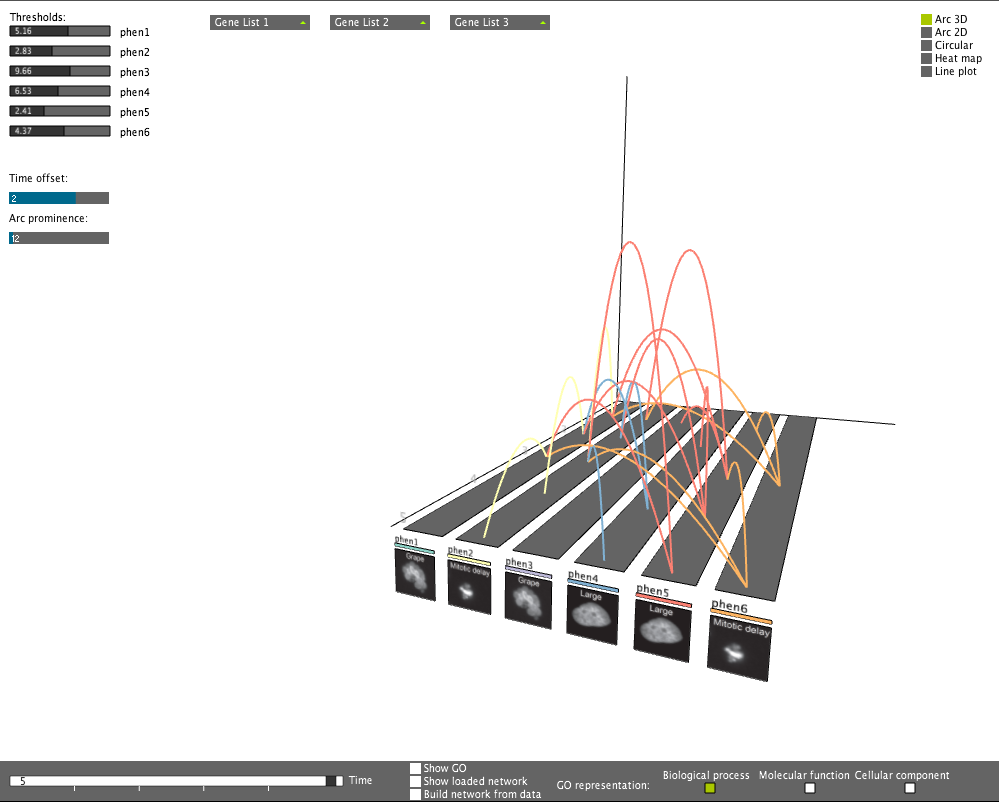 Setting phenotypic thresholds
Setting phenotypic thresholdsThresholds can be set for the phenotypes. Connections will be filtered correspondingly. Time offset can also be imposed.
-
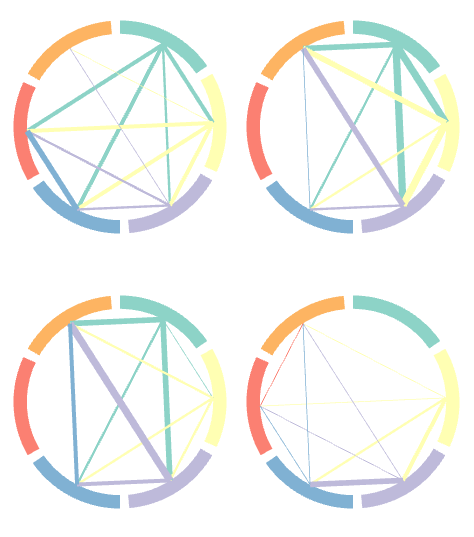 Phenotypic succesions in circular mode
Phenotypic succesions in circular modeCircular plots of phenotypic connections are drawn for every single time point.
-
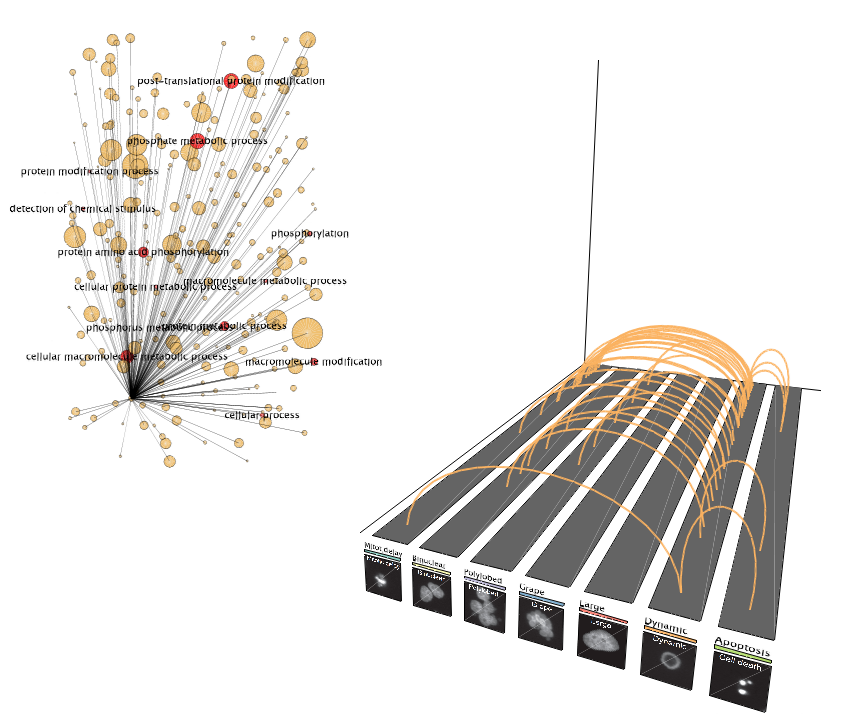 Single phenotype view
Single phenotype viewThe connections for one phenotype at a time can be analyzed.
-
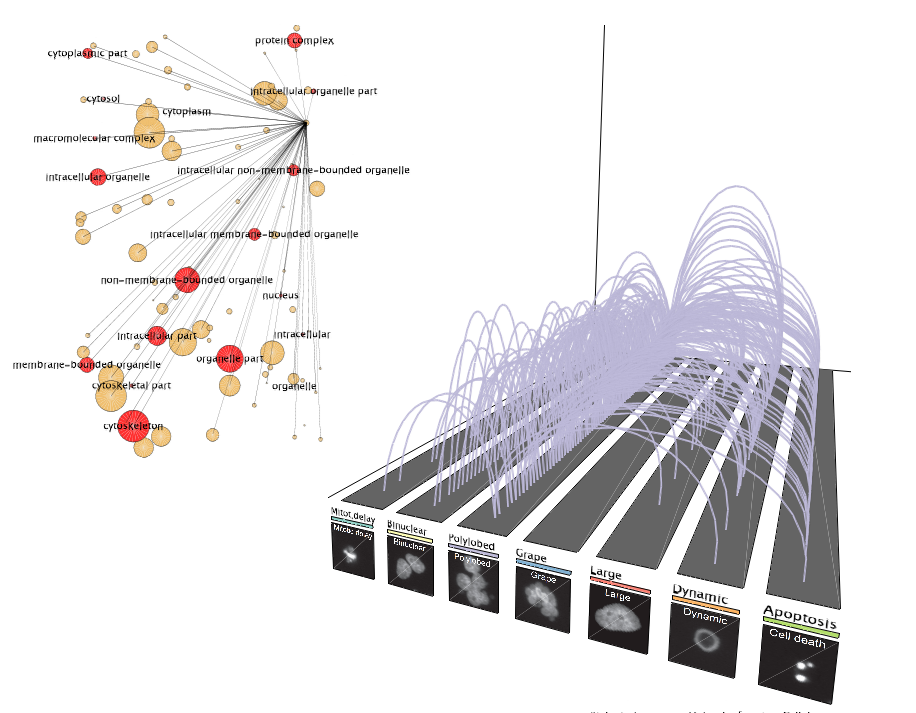 Polylobed transitions
Polylobed transitionsTransitions to the polylobed phenotype highlight interesting trends.
-
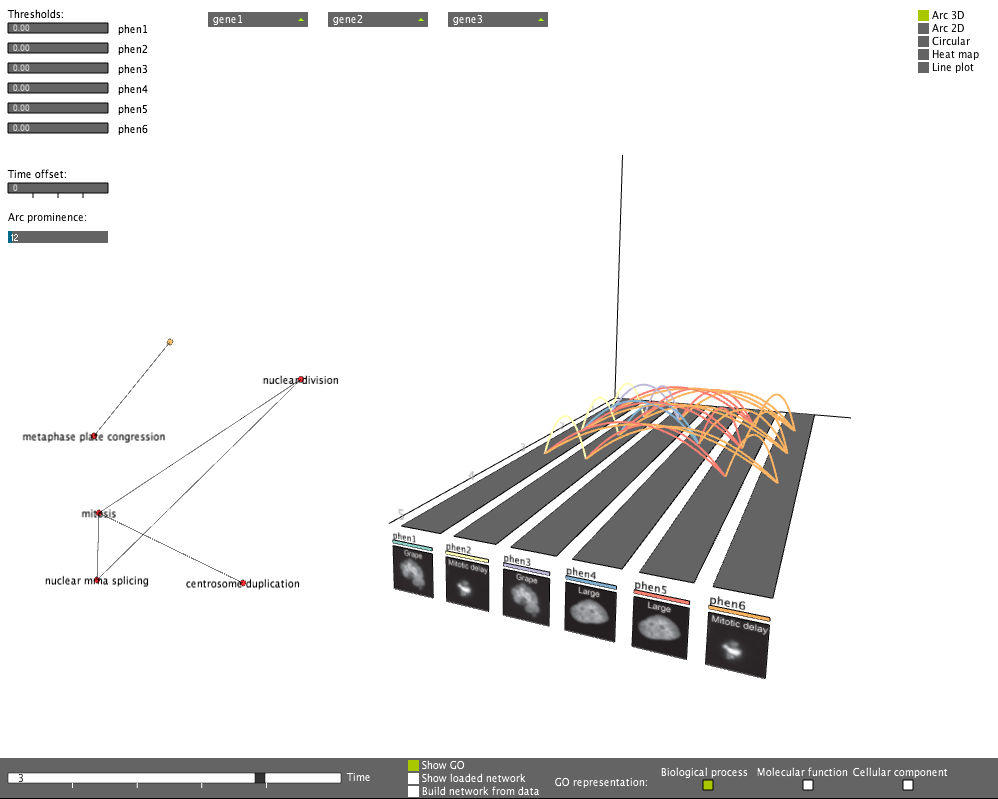 Dynamic GO network display
Dynamic GO network displayThe GO network highlights dynamically the enriched functionality of genes involved in connections at the particular time point.
-
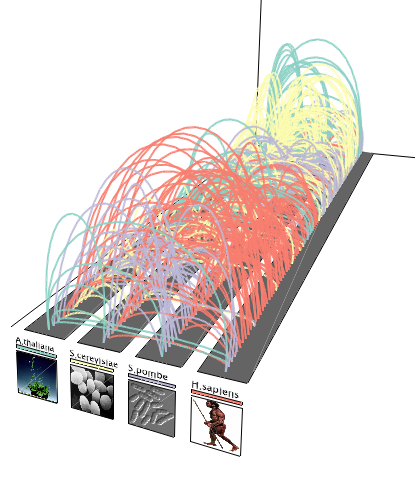 Orthologous links
Orthologous linksOrthologous cell cycle events are connected between four species.
-
Tool demo
This is a demo showing the main functionality of the tool. Warning: may not display well in Chrome browser.
