The different view modes
There are different visualization modes in PhenoTimer:
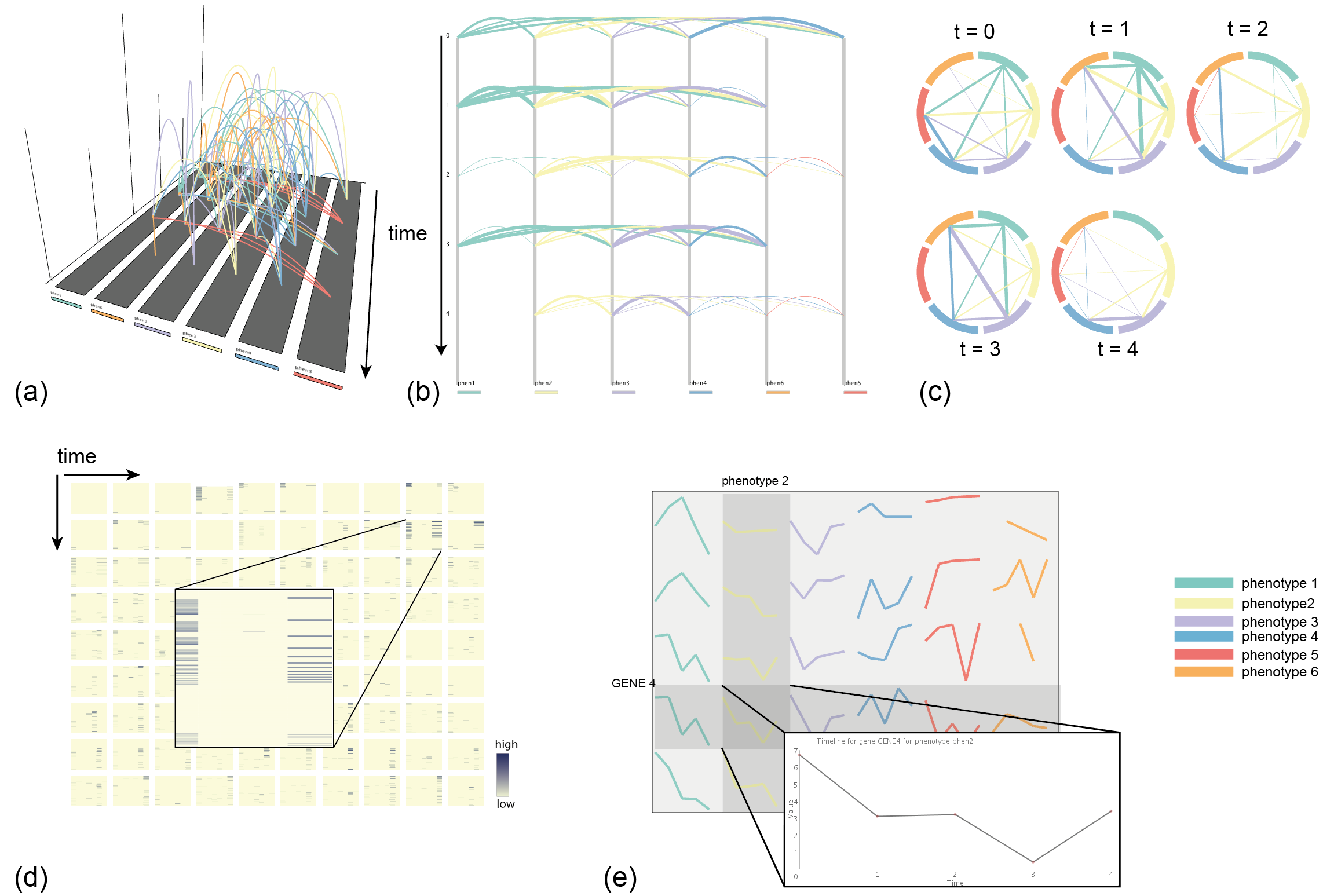
(a) 3D arc view (default)
Connections between phenotypes (or other variables) are represented as arcs, the color encoding directionality
(in case it is not needed, single color depictions can be used). The height of the arc corresponds to the number of genes
involved in the connection. Bar charts with values for every time point can be loaded as well.
(b) 2D arc view
Connections are represented as 2-dimensional arcs, with the same color coding as in (a). The thickness of the arc corresponds
in this case to the number of genes.
(c) Circular view
The connections between phenotypes are visualized in a circular manner for every time point.
(d) Heat map view
The gene-associated values are visualized for each phenotype in a separate heat map for every time point.
The rows and columns are clustered using agglomerative hierarchical clustering. The user can choose between single, average
or complete linkage and Euclidean or Manhattan distance.
The heat maps are expanded upon hovering and can be individually analyzed.
(e) Line plot view
The gene-associated values are visualized as timeline plots for every phenotype. The graphs are expanded upon mouse hovering.
